The Case of the Missing Prolines
Absolutes are red flags in microbiology
Microbiology is cool for many reasons, but bacterial diversity is one of the most exciting. Known bacterial diversity is constantly expanding. New techniques have taken us from 35 known phyla in 2002 to over 160 today.
The more we look, the more weird bacteria we find, and those discoveries keep overturning ideas that once seemed like laws.
We’ve seen complex internal organization within the PVC subphylum, multicellular behavior and differentiated cells in the Cyanobacteria, and even cases from my own work (shameless plug), where many bacteria don’t encode rRNA in the textbook fashion.
All this means that absolutes in microbiology are suspicious. If you try, you can often find some weird species that does x, y, or z differently than the established paradigm. What does this have to do with anything? LET ME TELL YOU.
An “invariant” feature across all domains of life
Around 2021 I was finishing up a comparative genomics project looking at amino acid sequence patterns across thousands of bacterial species. Specifically, polyproline sequences, where two or more prolines are encoded next to each other.
Polyproline sequences are interesting because they stall translating ribosomes. Proline is so structurally rigid that ribosomes have trouble incorporating it into new proteins. These aren’t rare sequences either—E. coli encodes over 2000 and humans encode more than 75,000. Ribosomal stalling at polyproline sequences is an intrinsic problem that all of life must deal with.
Proline, the trouble maker
Luckily, life is good at problem-solving! In bacteria, this issue is mainly solved by elongation factor P (EF-P). EF-P reduces the severity of ribosomal stalling at polyproline sequences by repositioning the proline carrying tRNA within the ribosome.
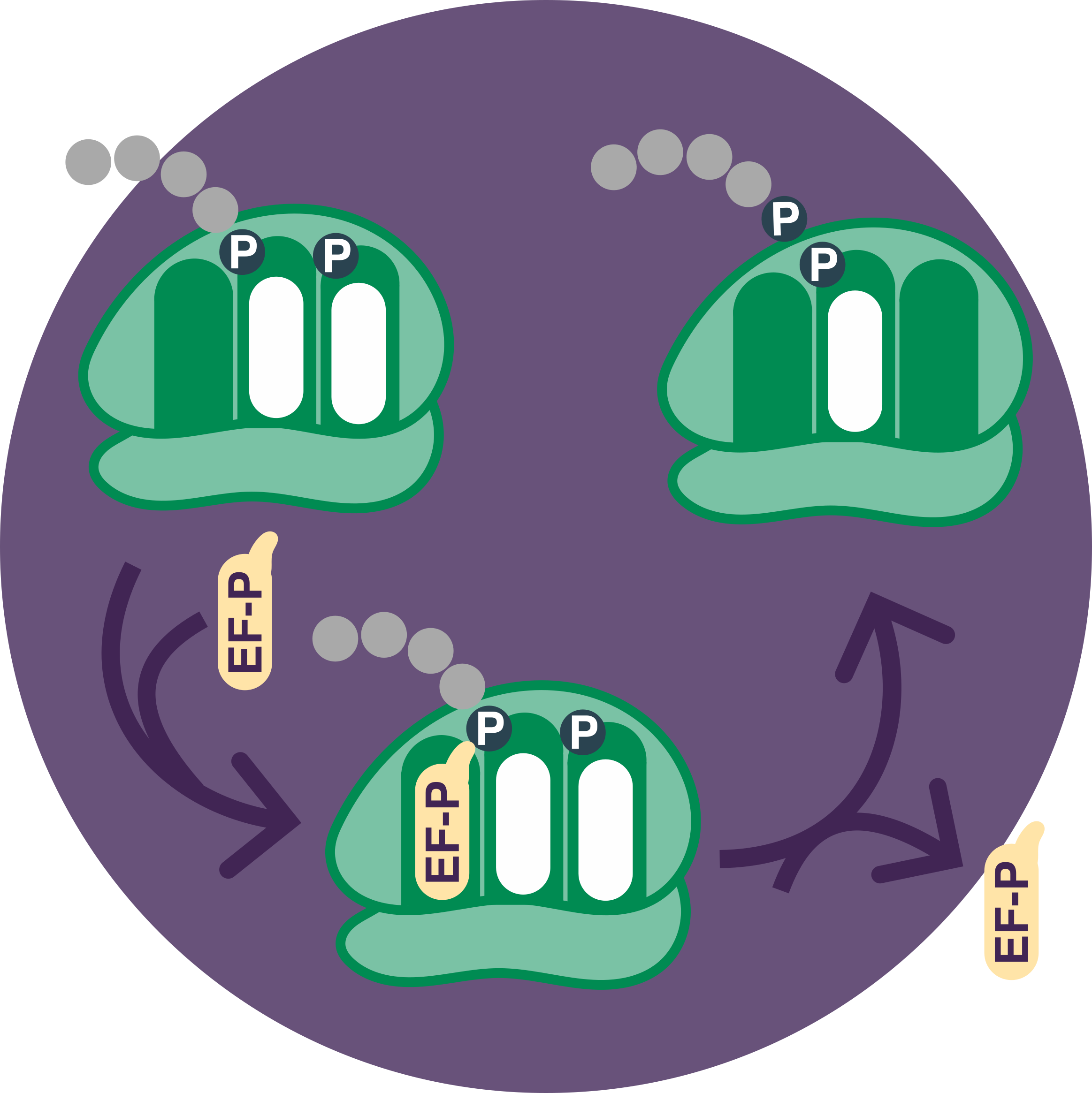
EF-P enters the ribosome and adjusts the position of tRNA to increase the rate of proline-proline bond formation.
Anyway, during this time, I was reading lots of literature on EF-P and polyproline sequences for the project. One paper (which despite what I write here is actually really cool and I like a lot) made a very bold claim:
“We have discovered that only a single polyproline stretch is invariant across all domains of life, namely, a proline triplet in ValS, the tRNA synthetase that charges tRNAVal with valine.”
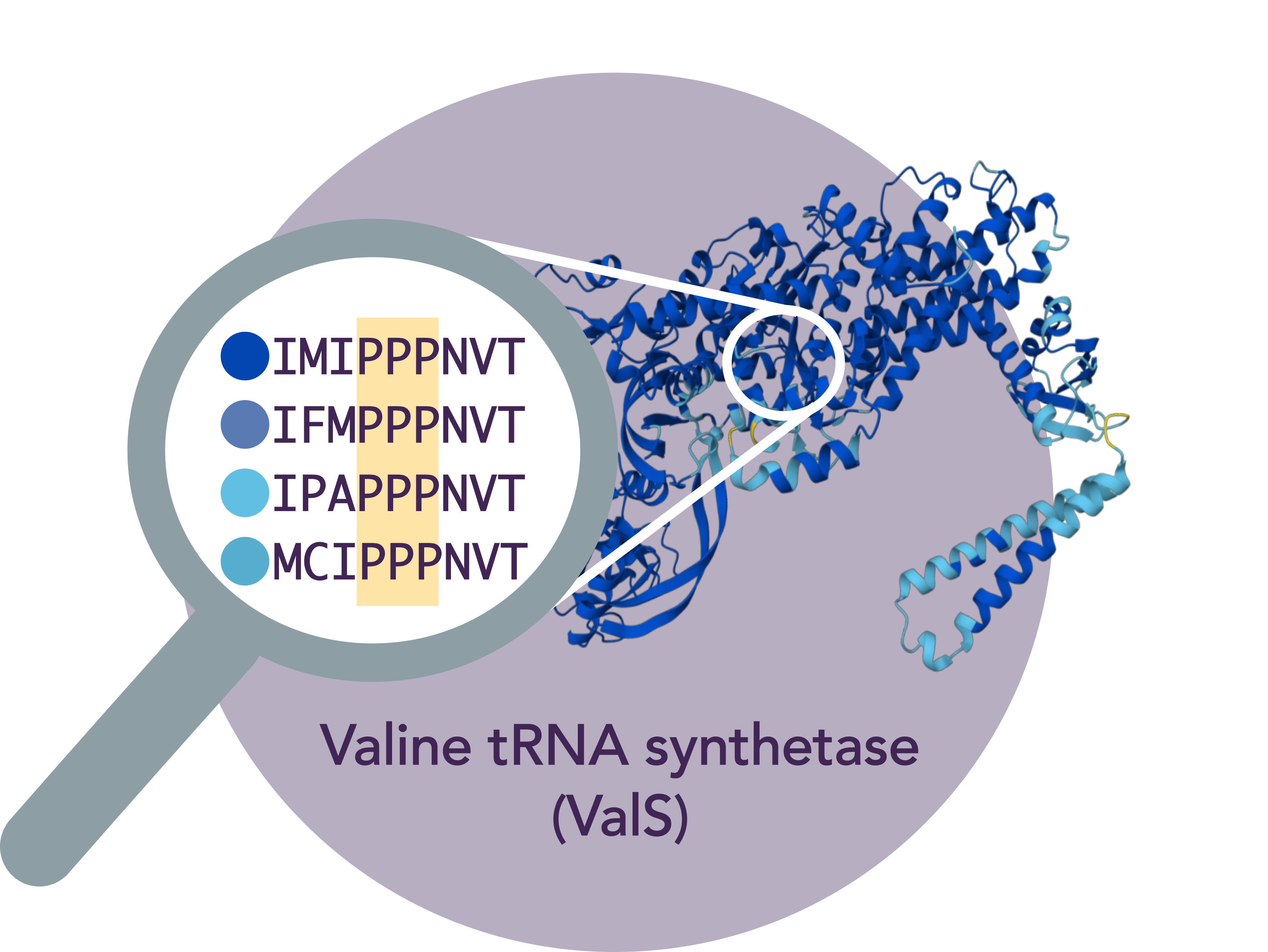
ValS sequences from top to bottom: E. coli, T. thermophilus, S. cerevisiae, H. sapiens
This statement piqued my interest, as all absolutes in microbiology do.
The exception that started it all
Since I already had a dataset of genomes where I had counted polyproline sequences, I decided to check this claim myself.
I found that almost every genome in my dataset fit this pattern, except one group. One family of Planctomycetes did not encode this “PPP” sequence. Instead, they encoded something slightly different: “PLP”. Upon closer inspection, these Planctomycetes also lacked other polyproline sequences that are conserved across most forms of life. What’s more, they had even lost entire proteins that normally contain polyproline sequences!
That was cool, but it got cooler. These same Planctomycetes did not have their usual version of EF-P. Instead, they had completely different versions that appeared to have been acquired from other bacteria, through horizontal gene transfer.
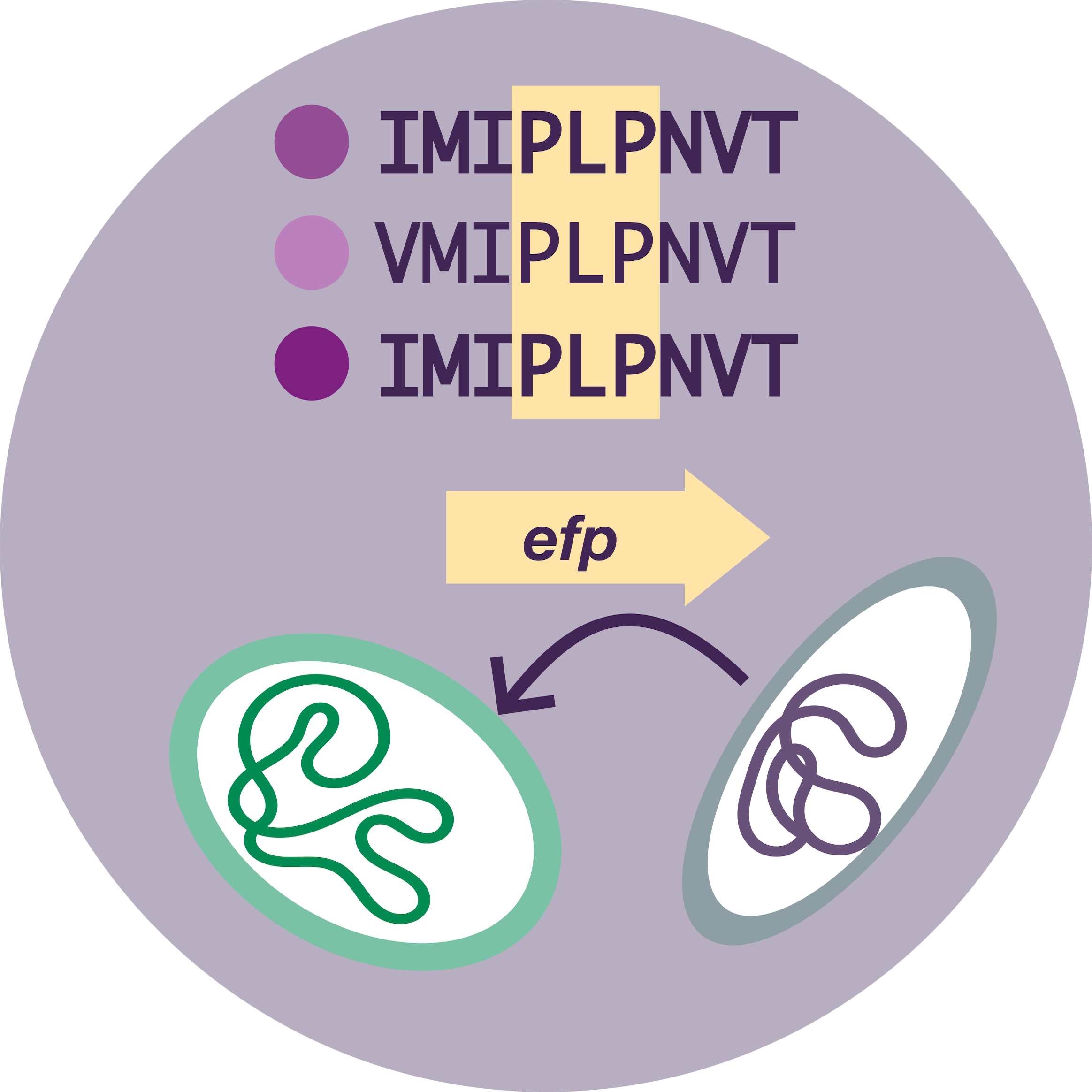
Planctomycete ValS sequences from top to bottom: P. limnophilia, A. californiensis, G. maris
This seemed like a pretty big coincidence and made me wonder…Was there was there a connection between the horizontal transfer of EF-P and polyproline loss?
A pattern consistent across distantly related bacteria
To find out, I looked at other groups of bacteria that have picked up foreign versions of EF-P. I found a similar pattern in a totally unrelated group, the Thermotogota. Once again, the appearance of a foreign EF-P co-occurred with the loss of highly conserved polyproline sequences and the proteins which contain them.
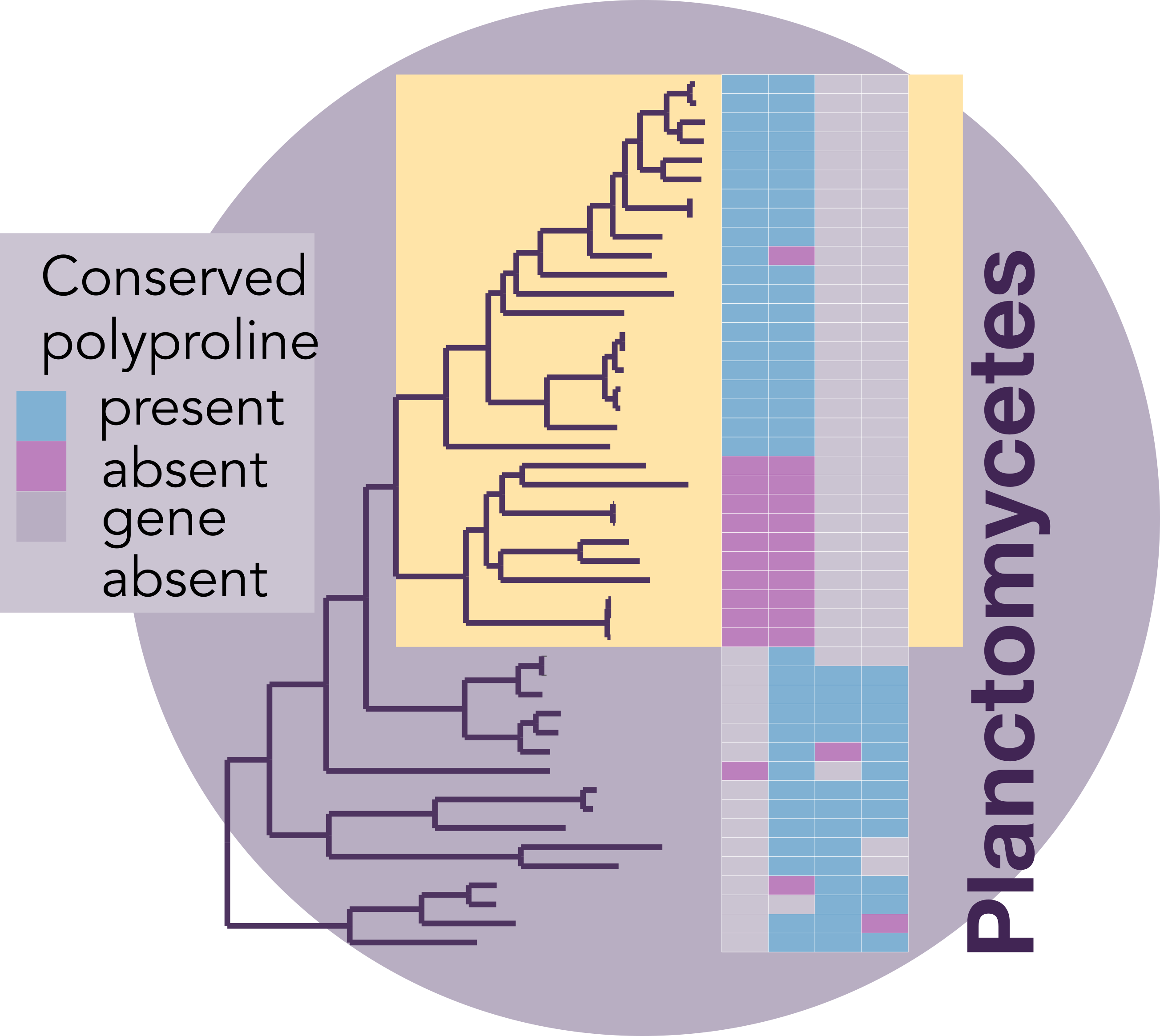
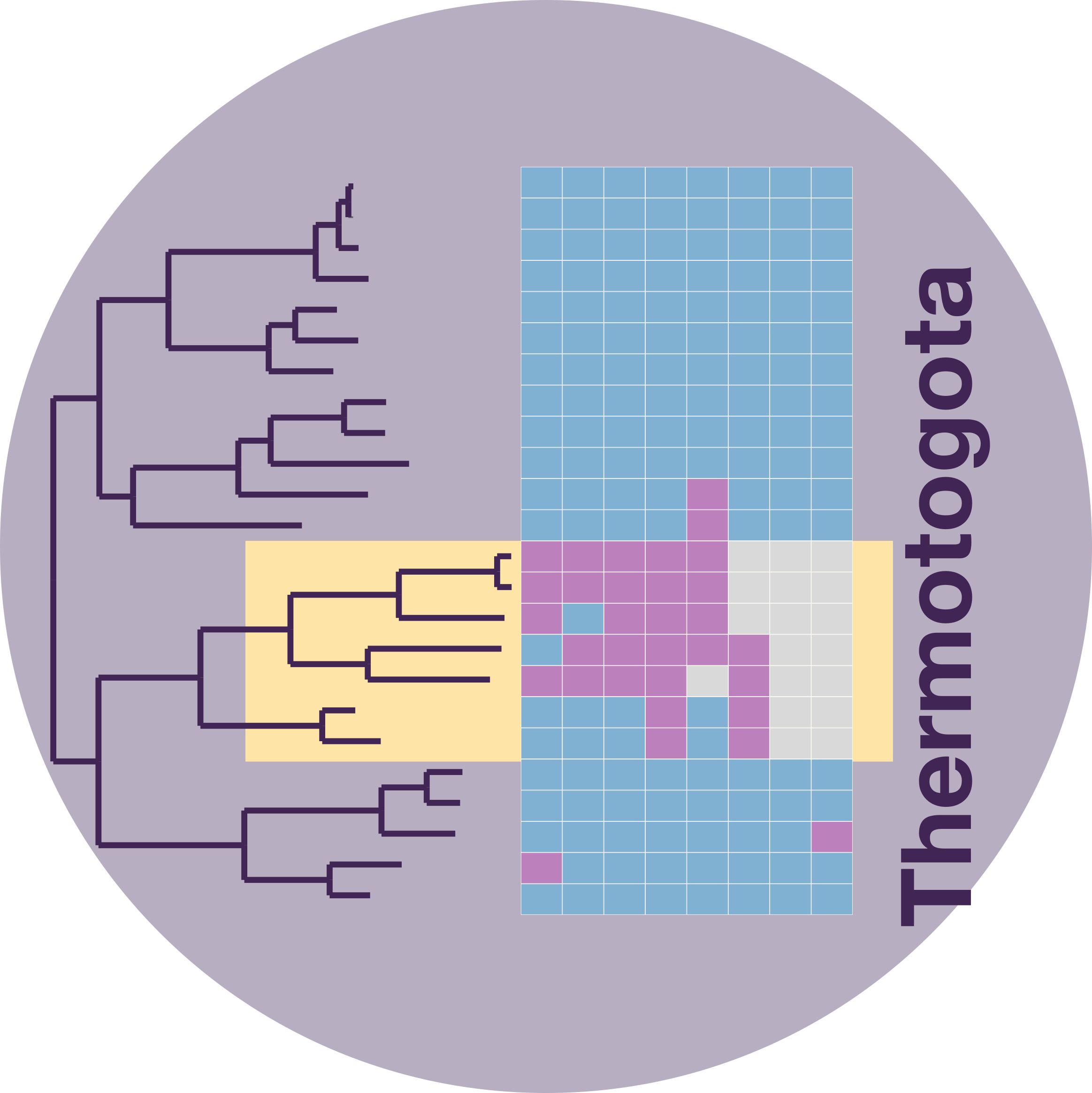
Phylogenetic tree where species with horizontally transferred efp are highlighted in yellow. Each column in the associated heatmap represents a protein where conserved polyproline sequences (purple) or entire proteins (grey) were lost. Proteins in the heatmap from left to right in are: (Planctomycetes) IleS2, ValS, IleS1, Lon and (Thermotogota) YcaJ, FtsH, ClpC, RpoD, TrpB, Lon, PilC, and TreT.
Two very distant groups of bacteria showing the same pattern suggested this wasn’t random. Something about acquiring a new EF-P system seemed to be linked to losing the very sequences that EF-P helps translate. But why?!
A hypothesis forms
To understand what might be going on, we first need to discuss a bit more about EF-P.
Many (but not all!) EF-P need to be chemically modified after they’re made in order to function properly. These modifications lengthen the part of the protein that reaches into the ribosome, and are thought to allow a better connection to the tRNA.
When these modified EF-P are horizontally transferred, they typically come together with the enzymes that attach the modification. For example, here’s a plasmid isolated from environmental microbial communities.
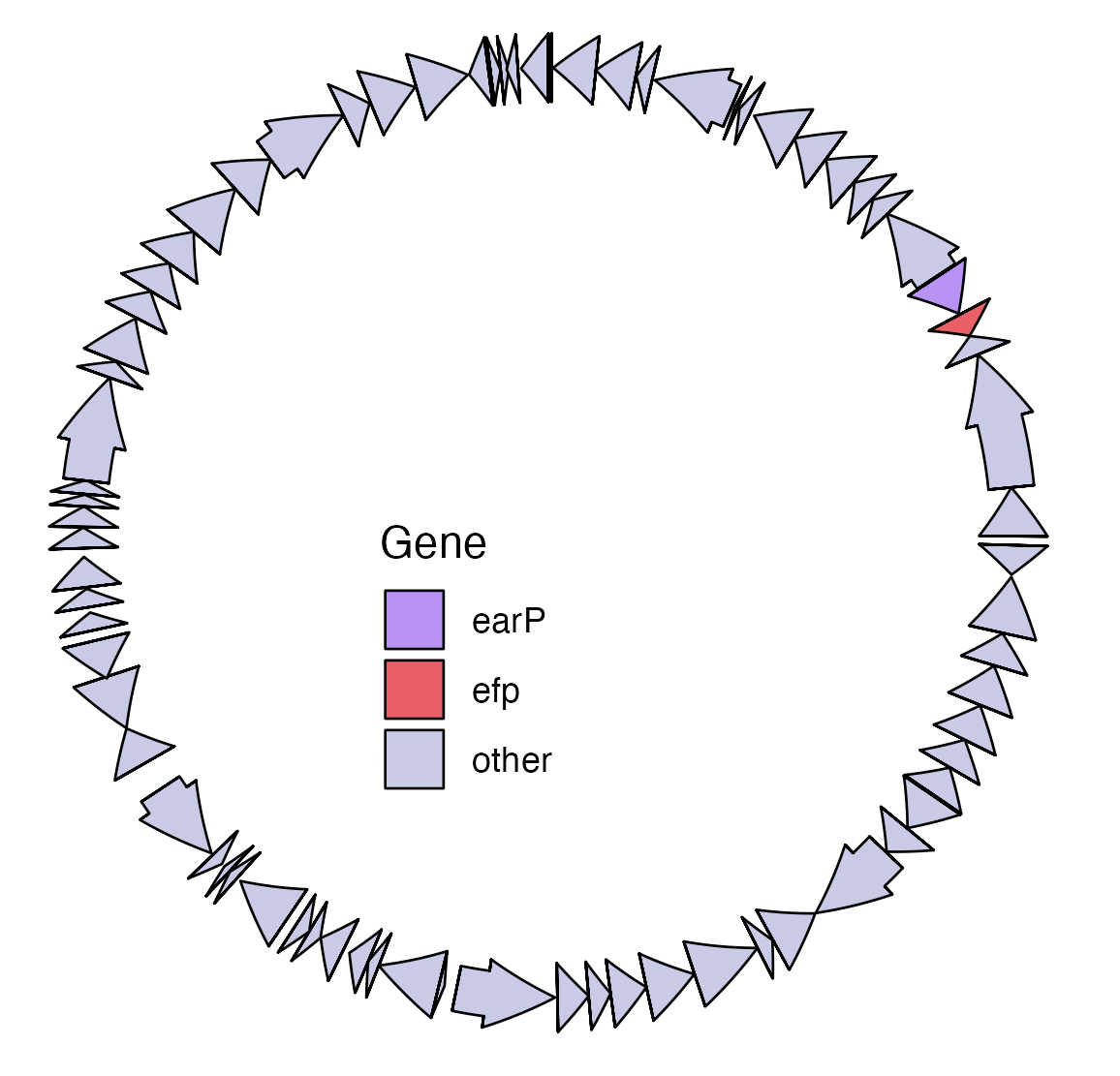
Environmental plasmid from desert spring benthic microbial communities showing co-location of an efp and earp gene. Each arrow represents an individual gene.
This plasmid encodes both:
- a modified EF-P, in this case with the sugar molecule rhamnose, and
- the enzyme that attaches this particular modification, EarP
My hypothesis, broken down sequentially goes like this:
- A bacterium acquires a new modified EF-P along with its modifying enzyme
- That enzyme doesn’t just modify the new EF-P; it also modifies the bacterium’s original EF-P
- The “unnatural” modification impairs the function of the original EF-P
- This reduces the cell’s ability to deal with polyproline sequences
- Over time, polyproline motifs are lost to make proteins easier to express
In short: importing a new system might accidentally break the old one.
Putting it to the test
My work is usually computational, analyzing genomic data. But to test my idea I really needed lab experiments.
So I joined a lab that specializes in studying EF-P, the Lassak lab at LMU Munich. Funnily enough, Jürgen Lassak is also the second author on the paper whose claim about an invariant polyproline sequence I had disproved. That was only slightly awkward.
Anyway, the plan was simple: I would take an EF-P that normally isn’t modified (I chose one from the Thermotogota Mesotoga prima) and expose it to a modifying enzyme (in this case EarP), and see what happens.
After quite a bit of work in the lab, the results were clear. EarP successfully added a rhamnose group to the “wrong” EF-P!
Even more importantly, this misplaced modification seemed to make the EF-P work less effectively. Experiments and modeling suggested that the added rhamnose physically interferes with how this EF-P interacts with the ribosome.
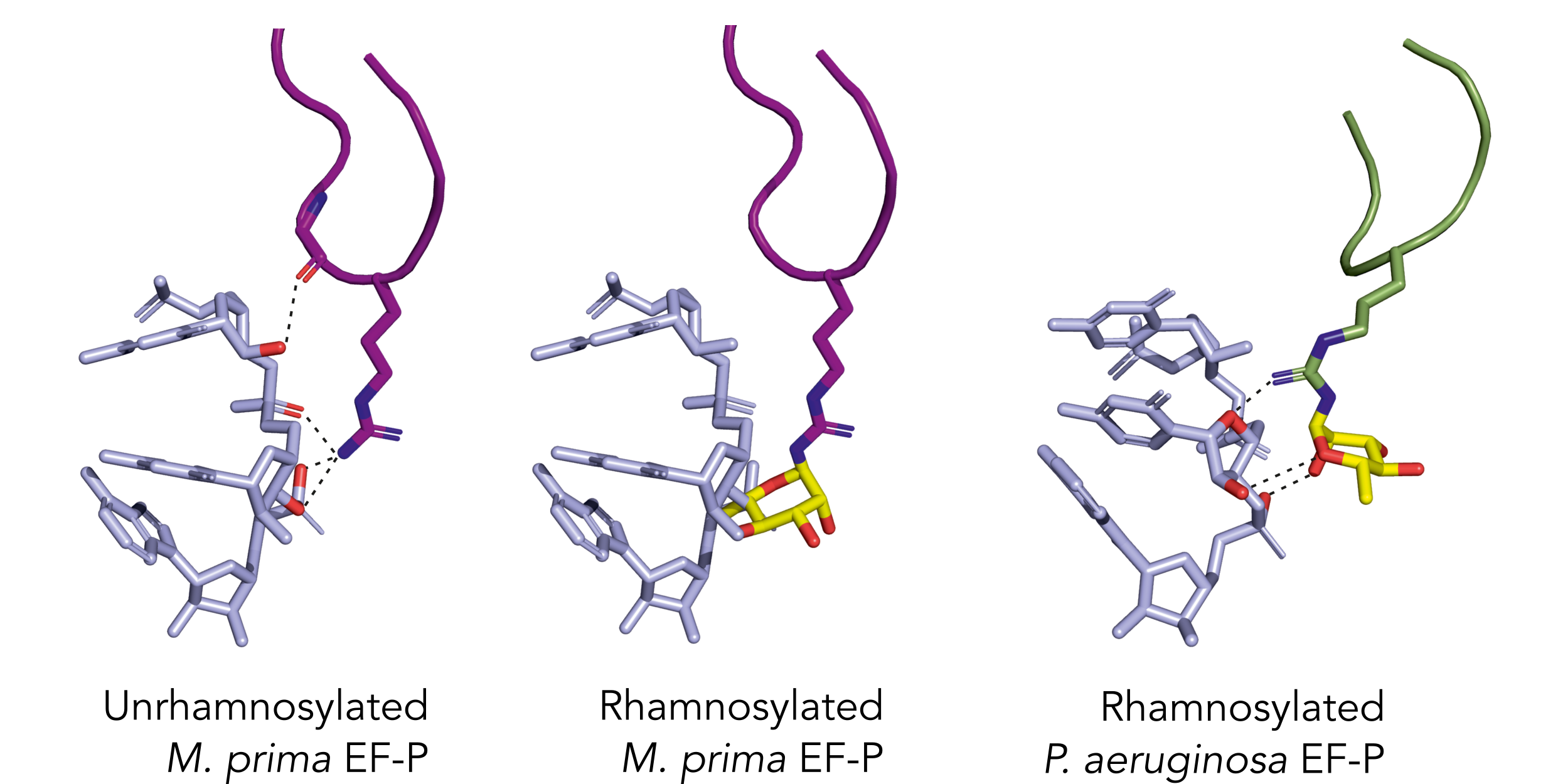
Models suggest that adding a rhamnose to the M. prima EF-P disrupts its function. Dashed lines highlight interactions between EF-P and the tRNA; rhamnose (in yellow) abolishes these contacts in M prima, unlike in P. aeruginosa, which is naturally rhamnosylated.
In other words, the new system was indeed disrupting the original one.
Take home – why bacterial diversity is important to study
Bacteria are constantly innovating, swapping genes, and reshaping their biology. What’s more, they’ve been doing this for billions of years! Their diversity is a massive archive of evolutionary history.
In this case, what started as a tiny exception to a supposed universal pattern led to a much bigger insight: when bacteria acquire new genes, the consequences can ripple through the cell in unexpected ways.
Horizontal gene transfer isn’t just about gaining new abilities, it can also create conflicts with existing systems, forcing the organism to adapt in response.
And perhaps most importantly, this story reinforces a core lesson in microbiology:
“Universal” rules are often just rules that haven’t broken yet.
This work was eventually published as two separate papers that are much more detailed than what I present here. If you’d like to check these out, here’s a link to the first and the second.